|
|
|
The lifetime of the signal transduced by Ras is determined by the life time
of the active state. If it is artificially prolonged, the biological response
may be unregulated and leads to drastic consequences in the cell. Ras was
originally found as the active principle of rodent tumor viruses and was called
an oncogene, a gene that is able to induce tumors in animals. Activated forms of
the Ras gene are found in up to 30% of human tumors. The activation of Ras is
due to a point mutation at either position 12, 13 or 61, the biochemical
consequence is to render the protein unable to hydrolyze GTP. Since many other
genes involved in the Ras signal transduction pathway are also found as
oncogenes in human or animal tumors, Ras itself and the Ras pathway is thus
considered to be a prime target for anti-tumor therapy.
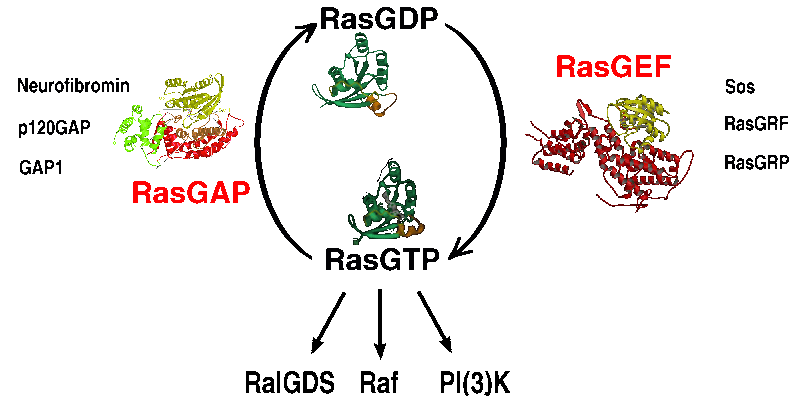 Ras is switched ON by several GEFs, the most important of which are Sos
(derived from Son-of-sevenless), Ras-GRF and Ras-GRP. In the activated GTP-bound
state it interacts with several downstream target proteins such as the protein
kinase Raf (of which three isoforms c- , A- and B-Raf exist),
Phosphoinositide-3'-kinase (=PI(3)kinase, 4 isoforms) and Ral-GDS (at least four
isoforms (RalGDS, Rgl, Rgr, Rlf). Each of these effectors is believed to
contribute to the Ras signal. The best understood of these signals is the
activation of the Raf kinase which initiates the activation of the Map kinase
Erk via the Map kinase kinase MEK. Ras-GTP is switched OFF by the action of
specific Ras-GAPs, the most studied of which are p120GAP and neurofibromin, the
gene product of a tumor suppressor gene.
Ras is in green, the GAP in red color. The nucleotide GDP and the aluminum
fluoride is shown with white bonds, and you can see the "finger loop" of the GAP
protruding into the nucleotide binding pocket of Ras. A large part of the Ras
protein contacts the GAP which stabilizes some flexible regions of Ras which are
important for the nucleotide hydrolysis reaction.
This movie shows the conformational changes of Ras upon hydrolysis of GTP to GDP. The "effector loop" or "switch I region" is shown in red, the second region which shows major changes - the "switch II region" is highlighted in cyan. The gamma phosphate group determines the structure of the flexible switch regions. (download
as MPEG movie, ca. 0.4 MByte)
The GTP-binding protein Rap is a close homologue of Ras, with
50% identity in amino acid sequence. There are four isoforms, Rap1A and B, and
Rap2A and B, with Rap1 less divergent from Ras then Rap2. Rap1 has originally
been isolated as a suppressor (revertant) of K-Ras transformation (hence its
other name K-rev-1), and was shown to inhibit the Ras-Map kinase pathway.
Nowadays it is believed to have an independent biological role, most likely that
of activating integrin, the cell surface protein involved in cell-cell
interactions. 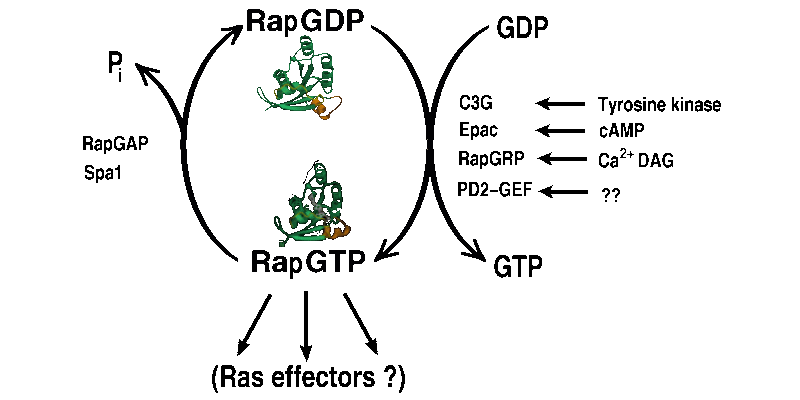
The Ras-binding-domain (RBD) of the Raf kinase is shown in red, Rap is in
green and the nucleotide is depicted as a ball-and-stick model. The interaction
accurs mainly via the two adjacent beta sheets of Raf and Rap. Groups working on Rap: AG A. Wittinghofer , AG R. Ahmadian , AG C. Herrmann
Rho GTP-binding proteins regulate a variety of cellular
processes in all eukaryotic cells through the reorganization of the actin
cytoskeleton resulting in cytokinesis, phagocytosis, pinocytosis, morphogenesis,
axon guidance, cell polarity, cell migration. To date 16 different mammalian Rho
GTPases have been identified which are members of the Ras superfamily. The most
extensively characterized members are RhoA, Rac1 and Cdc42.
The GTPase-cycle is regulated by numerous cellular proteins. More than 30
Rho-specific GEFs are known all of which contain a Dbl-homology (DH) domain
responsible for the catalytic activity and an adjacent pleckstrin homology (PH)
domain. Around 30 potential effectors for the Rho-family GTPases have been
identified which are Ser/Thr-Kinases (Pak, Rock) or function as scaffold
proteins (WASP, mDia). In addition about 20 Rho-specific GAPs are known.
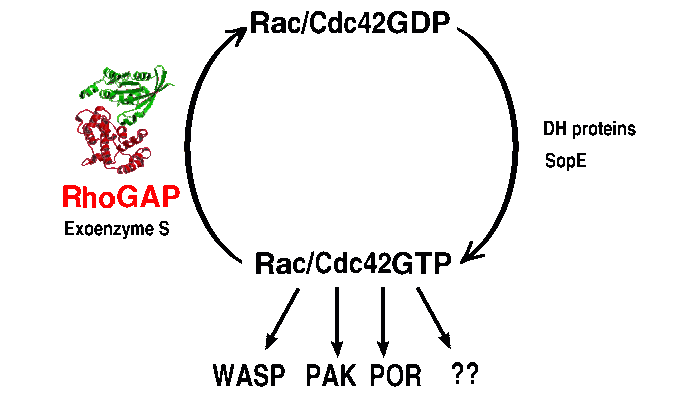 Groups working on Rho/Rac/Cdc42:
Ran (the Ras-related nuclear protein) is the major regulator of
nucleo-cytoplasmic transport across the nuclear pore complex (NPC). Like other
small Ras-like GTP-binding proteins it switches between a GTP- and a GDP-bound
form by GTP-hydrolysis and nucleotide exchange. In contrast to its relatives Ran
does not undergo posttranslational modification. The exclusive nuclear localization of the Ran exchange factor RCC1 ensures
that the generation of Ran· GTP is confined to the
nucleus. The conversion of Ran· GTP into Ran· GDP is catalyzed by the GTPase activating protein RanGAP1,
which is present only in the cytoplasm or at the cytoplasmic site of the
nucleare pore, efficently depleting Ran· GTP from the
cytoplasm. This differential localization of the regulators of Ran's
nucleotide-bound state should thus result in a Ran·
GTP-gradient across the NPC which is believed to drive the import-export cycle.

Ran interacts with transport receptors which load and unload their cargo in the respective compartment depending on the nucleotide state of Ran. An importin binds its cargo initially in the cytoplasm, gets translocated through the NPC, releases the cargo upon binding Ran· GTP in the nucleus and returns to the cytoplasm as a Ran· GTP complex. The removal of Ran· GTP from the importin involves the hydrolysis of the Ran-bound GTP and allows the importin to bind and import the next cargo molecule.
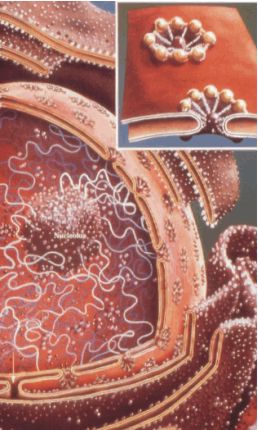
Fig. A: The nuclear pore complex (NPC). The nuclear envelope separates cytosol and nucleus and guaranties efficient
separation between genetic information and protein synthesis or metabolic
processes. Transport between nucleus and cytoplasm is organized by nuclear pore
complexes which are huge complexes of approx. 125 MDa assembled by 100-200
different proteins. Every minute several million of protein and RNA-molecules
must be transported between the cytoplasm and the nucleus of a eukaryotic cell
and have to pass the nuclear pores.
Binding of substrates to exportins is regulated in a converse manner to importins. Exportins bind their cargos preferentially in the nucleus, forming a trimeric cargo· exportin· Ran· GTP complex. This trimeric complex is then transferred to the cytoplasm where Ran· GTP is converted to Ran· GDP. This results in Ran's dissociation from the complex, allowing the exportin to release its substrate, re-enter the nucleus, and to start the next export cycle. The recycling process of Ran between its GDP and GTP complexed state is supported by another class of Ran-binding proteins, which support GTP-hydrolysis by RanGAP and are either soluble proteins or part of the nuclear pore complex. Sequences of Ran and Ran-interacting proteins are highly conserved between different species and allow interspecies exchange of GEFs and GAPs from e.g. yeast to mammalian. Disturbing the Ran-system causes analogous effects on the organisation of the nuclear skeleton, RNA-processing and transport, protein import and cell-cycle control. 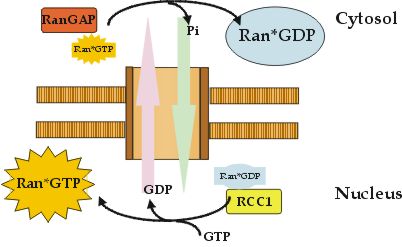 Fig. B: The cellular compartment defines the nucleotide state of Ran.
In the cytosol a Ran specific GTPase activating protein (RanGAP) ensures that Ran exists almost exclusively in the GDP-complexed state. In the nucleus a Ran nucleotide exchange factor (RCC1) catalyzes the exchange reaction from Ran· GDP towards Ran· GTP. Import and export complexes recognize their actual cellular environment by interaction with the Ran protein.
Several researchers of our department - partly in collaboration with other institutes - have contributed to information upon:
Our actual interest is focused on:
Groups working on Ran:
The human guanylate-binding protein 1 (hGBP1) belongs to the family of large GTPases. Its synthesis in cells is induced after activation by interferon-gamma. So far the biological function of hGBP1 is not known albeit an antiviral effect was demonstrated. Whereas there is little primary sequence homology to the other large GTPases like dynamin the relationship between the proteins becomes evident by the common architecture of the protein domains and by common biochemical features like nucleotide-dependent oligomerisation and cooperative GTPase activity.
|
|